We study the evolution of species interactions using interdisciplinary approaches. Each individual project has a specific focus that aims to address a major evolutionary question.
We study the evolution of species interactions using interdisciplinary approaches. Each individual project has a specific focus that aims to address a major evolutionary question.
In addition to these ongoing and funded projects, we are also working on developing new tools and resources in the area of evolutionary single-cell genomics.
Plant perspective
MapNTSR

Adaptive evolution of plants to environmental change – either natural or human-mediated - shaped the global biodiversity and can profoundly affect the development of human societies. A classic example is the rapid increase of herbicide resistance in weeds, which threatens worldwide crop yields. The mechanisms of herbicide resistance mechanisms are classified into two types: target-site resistance (TSR), which is caused by changes in the herbicide target protein, and non-target site resistance (NTSR), which includes all other mechanisms. Recent studies suggested that NTSRs arise from standing genetic variations that are associated with adaptation to natural stresses. However, due to the lack of a suitable model system in which the detailed molecular mechanisms and ecological functions of NTSR can be characterized, this hypothesis remains largely unexamined.
Here, we aim to address this challenge using S. polyrhiza as a model system.
EvolTroph
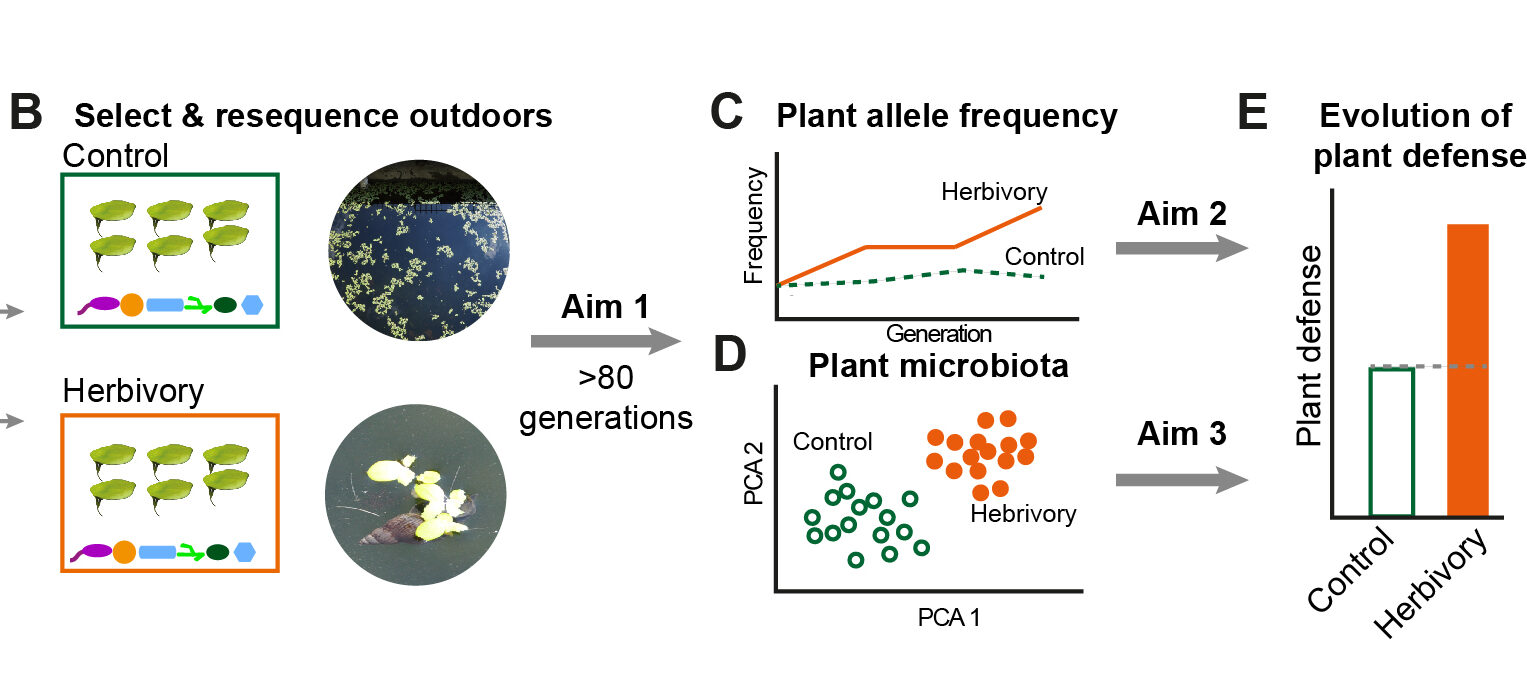
In nature, plants are threatened by herbivores and colonized by diverse microorganisms. Accumulating evidence suggests that the plant microbiota changes the host’s defense against herbivores and is altered by herbivory. However, it remains unknown to what extent interactions between plants and their microbiota affect the process and trajectory of plant evolution under herbivore pressure.
This project aims to close this knowledge gap by investigating the process and mechanism of herbivory-driven plant evolution in real-time. We will perform “selection and resequence” experiments using the giant duckweed (Spirodela polyrhiza), one of the fastest growing plants, as a model system. We will impose herbivory pressure on S. polyrhiza populations that represent the majority of genetic diversity in this species for more than 40 generations. We will quantify the extent to what herbivory drives changes in the plant phenotype, allele frequency, and microbiota. We will further assess the herbivory-dependent fitness consequences of the observed changes in plant traits and allele frequencies, and investigate whether and how changes in the plant microbiota alter the herbivory-imposed selection on plant defenses.
Duckweed evolutionary genomics

We are currently developing pan-genome resources for S. polyrhiza.
Herbivore perspective
EvolToxin

Many insects produce toxins in their hemolymph that are lethal to their predators but not to themselves. To achieve this, insects must specifically regulate the production of the toxins and evolve self-resistance at the cellular level. However, the underlying mechanisms and evolutionary processes remain largely unknown.
Leptinotarsa hemolymph produces a high abundance of leptinotarsin, a presynaptic neurotoxin that is lethal to many insects and vertebrates but harmless for the carriers (2, 3). The peptide suggests that the toxin evolved from a duplicated juvenile hormone esterase (JHE), which is specifically expressed in hemolymph (4). While the regulation of JHE is age-dependent (5), leptinotarsin is found in all developmental stages.
Our preliminary data suggest that leptinotarsin is produced in a specific type of hemocyte cells, suggesting the evolution of new gene regulations of leptinotarsin. Here, we will use tools from comparative genomics and single-cell sequencing to study the mechanisms and evolution of cell-specific regulations of leptinotarsin. In addition, we will investigate the mechanisms of toxin self-resistance using a functional neurobiology approach. Specifically, we will address three questions: 1) Which cells produce leptinotarsin and how is it regulated at the cellular level? 2) What are the mechanisms of the leptinotarsin self-resistance? 3) How did leptinotarsin and its resistance mechanism evolve? We will first perform single-cell RNA-sequencing (scRNA-seq) on Colorado potato beetle (Leptinotarsa decemlineata) hemocytes to characterize the cellular regulations of leptinotarsin (Aim 1). We will further use single-cell ATAC-sequencing to investigate how chromatin structures contribute to the evolution of cell-specific regulations of leptinotarsin. Second, we will investigate possible mechanisms of leptinotarsin self-resistance by investigating synaptic targets of leptinotarsin in Drosophila (Aim 2). Third, we will retrace the evolutionary history of leptinotarsin and the toxin self-resistance mechanisms by analyzing the genomes of closely related Leptinotarsa species and other insects (Aim 3). Comparing the target molecules in species that are susceptible to the toxin with resistant species will provide insights into the evolution of resistance mechanisms. We hypothesize that leptinotarsin evolved new regulations via changing cell-specific chromatin accessibility, the process of which is after the emergence of self-tolerance.
Together, this project will provide new insights into the evolution of novel gene regulatory networks at the single-cell level.
This project is in collaboration with Prof. Marion Silies and Dr. Claudia Keller Valsecchi.
This project is part of the GenEvo initiative.
EvolPest

Resistance to insecticides in insect pests is a remarkable example of rapid adaptation in a human-mediated evolutionary experiment. It has become apparent that insecticide resistance is unavoidable under current agricultural practices and threatens global food supply, highlighting the urgent need to investigate the evolutionary processes of insecticide resistance.
The Colorado potato beetle (CPB, Leptinotarsa decemlineata), an insect with little practical significance to crops 150 years ago, is now a devastating global pest of potato and other solanaceous crops and has evolved resistance to nearly all known insecticides. The remarkable resistance to insecticides and relatively short and well-known natural history thus make CPB an ideal system to study the mechanisms of insecticide resistance and evolutionary processes of agricultural pests. Here, we will investigate the evolution of CPB and its insecticide resistance by measuring key population genetic parameters and reconstruct the demographic history of CPB. Specifically, we will 1) estimate spontaneous mutation rates in CPB and evaluate the extent to which the mutation rates are affected by a low-dosage of insecticide in mutation accumulation experiments; 2) map genetic basis of resistance to two widely used insecticides through bulk-segregation analysis; 3) characterize the evolutionary history of CPB and its resistance by re-sequencing the genomes of globally distributed CPB populations.
The outcomes of this project will push the boundaries of our current knowledge of insecticide resistance evolution by providing both mechanistic and genomic insights into how current agricultural practices might have contributed to the evolution of a devastating pest. The obtained population genetic parameters may further facilitate the development of sustainable pest control strategies.
Niche choice & adaptive evolution
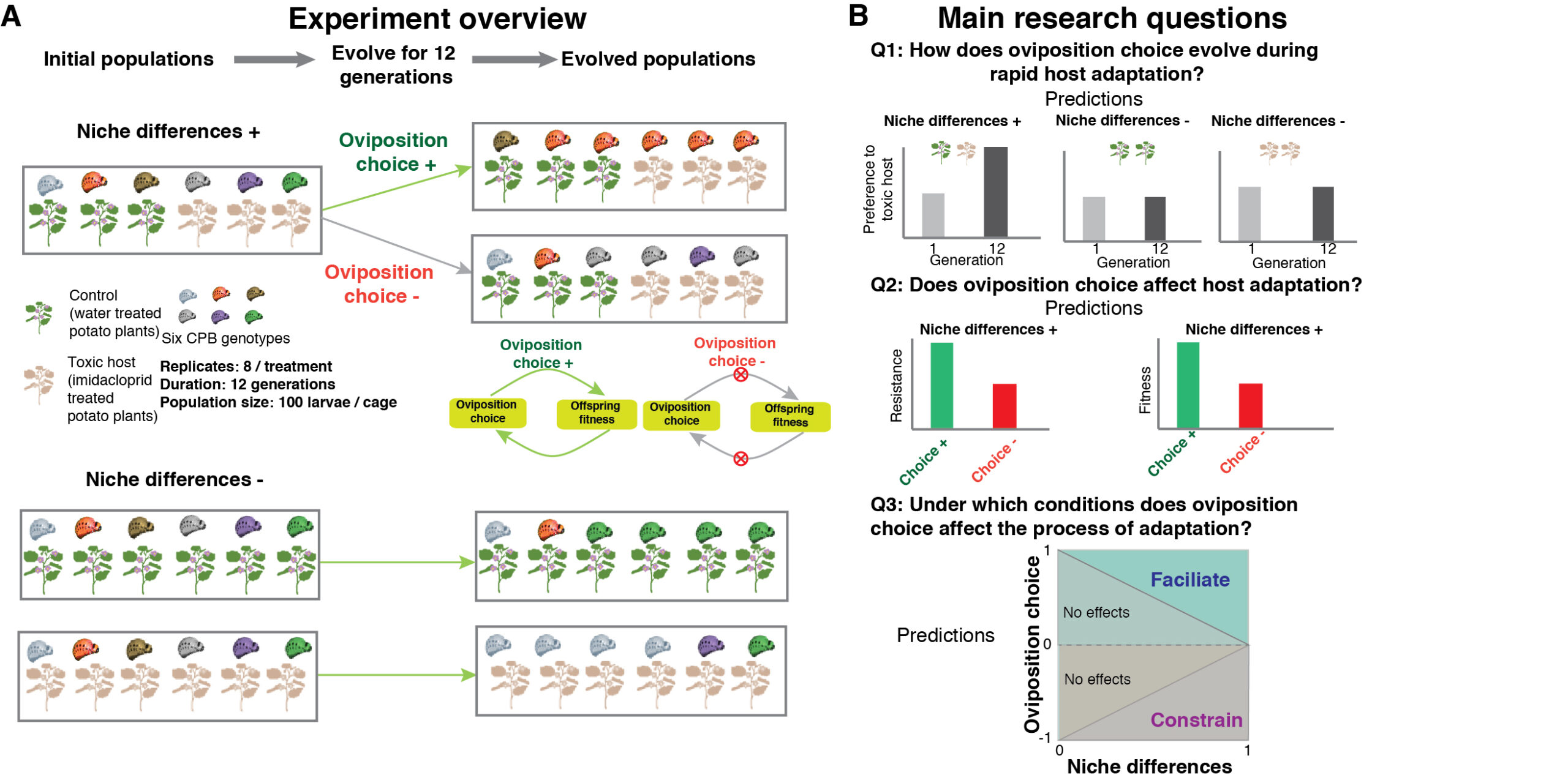
In this project, we will use an experimental evolution approach in combination with trait manipulation and computational modelling to investigate whether and how individualised niche choice affects the evolutionary process of host plant adaptation in Colorado potato beetles (Leptinotarsa decemlineata, CPB). This project will allow us to directly examine several long-standing theoretical predictions in niche concepts.